 A tool for designing gene-specific primers targeted at genomic regions of interests based on known gene structure and EST sequence information
A tool for designing gene-specific primers targeted at genomic regions of interests based on known gene structure and EST sequence information
The Expeditor is a web-based tool to combine human intron/exon structural information for a given gene and EST sequence information of the target species (or a good consensus coding sequence from multiple species), to design primers for amplification of genomic segments of a gene.
|
|
This software is free for use by not-for-profit or academic users. However the users are strongly encouraged to send us feedbacks, which is important for improvements.
The Expeditor is made intuitively easy to use (See our Three-Step-Guide). One can simply follow instructions at each step as you go through the designing process on this web tool.
Some 3rd party software / rules are embeded in Expeditor. We want to acknowldge the following:
- BLAST Program:
Used to find best sequence equivalents of each exon from your targeted species.
- Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman, (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs, Nucleic Acids Res. 25:3389-3402.
- CAP3 Program:
Used to align several sequences from different species for their consensus.
- Huang, X. and Madan, A. (1999) CAP3: A DNA Sequence Assembly Program, Genome Research, 9: 868-877.
- Single letter nucleotide symbol (IUPA code):
Used to indicate non-consensus nucleiotides.
- Nomenclature Committee of the International Union of Biochemistry (NC-IUB) (1984) "Nomenclature for Incompletely Specified Bases in Nucleic Acid Sequences"
- Primer3 Program:
Used to design primers on the template sequence formed.
- Rozen, S., Skaletsky, H. "Primer3 on the WWW for general users and for biologist programmers." In S. Krawetz and S. Misener, eds. Bioinformatics Methods and Protocols in the series Methods in Molecular Biology. Humana Press, Totowa, NJ, 2000, pages 365-386.
This tool has been published on Journal of Heredity, January 2005 issue. Users may cite the paper as:
Hu, Zhi-Liang, Kimberly Glenn, Antonio M. Ramos, Charles J. Otieno, James M. Reecy and Max F. Rothschild (2005) Expeditor: A Pipeline for Designing Primers Using Human Gene Structure and Livestock Animal EST Information. Journal of Heredity, Volume 96, Number 1 (January 2005), pp. 80-82
| Google Scholar citations: 15 times | 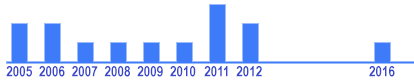
|
This work is supported by funds from the USDA National Animal Genome Research Program, Pig Genome Coordination and Bioinformatics Coordination Programs.
|
|